A novel mechanism of combating inherent DNA replication problems?
By Syed Moiz Ahmed, PhD Student, Nanyang Technological University
Cancer cells, like other fast proliferating cells, undergo DNA replication at overwhelming speeds, which puts them in a precarious situation with respect to DNA topology. Excess underwound or overwound DNA can lead to premature termination of replication. DNA supercoiling (the coiling of the DNA about itself) has been an often-overlooked, unappreciated biological process, but it is rising to prominence as a force that drives life as we know of it today.
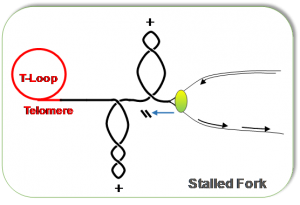
Figure depicting DNA structures such as three-way junctions, single/double strand DNA and supercoiled DNA at stalled forks.
In response, all cells have come up with tools to effectively counter and run the engines of replication smoothly. Although several of these have been widely studied, such as DNA damage response and repair systems, none seem to be safe and economical. A profitable and more logical approach for cancer cells would be to use more direct and energy-efficient processes that work at the root of the problem.
To study this, a research group at Nanyang Technological University in Singapore has found an oncofetal, transcription factor called HMGA2 that co-localizes with key replication factors and prevents the collapse of stalled replication forks. The paper published in Cell Reports shows HMGA2 uses all three of its AT-hook motifs to bind to DNA and prevents the formation of pathological structures at the fork that are caused due to replication stalling.
In a more recent study from the same group published in Nature Scientific Reports, HMGA2 was found to bind with high affinity to supercoiled DNA. This novel mode of DNA binding and scrunching discovered through single molecule and atomic force microscopy techniques advances and opens up our understanding of topological regulation that could occur in cancer cells.
Some key questions yet remain. Topoisomerases, enzymes that catalyze the release of supercoiling at replication forks, are very well studied and are prominent chemotherapeutic targets of several cancers. However, cancer cells can astonishingly cope with these topological challenges thrown at them, leading to failed therapeutic responses in patients. One would wonder, how does this work?
To answer exactly this, the team plans to continue studying the regulation of DNA topology in context of the oncofetal HMGA2 protein and by targeting topoisomerases. They not only hope to unravel mechanisms that bring out a clearer picture on replication stress regulation but also be able to reveal potential therapeutic targets.
Syed Moiz Ahmed is a PhD Student at Nanyang Technological University in Singapore.
If you’re interested in writing for our blog, get in touch with us at info@kerafast.com. Articles submitted by July 31, 2018 will be entered in our summer blog contest for a chance to win a $500 Amazon gift card! Contest rules can be found here.


