New details on antibiotic resistant bacteria
Bacteria gain resistance to antibiotics via mutation events in their genome. A recent study published in Nature Communications sheds new light on a gene that decreases mutation rates in Mycobacterium smegmatis, a model for Mycobacterium tuberculosis.
M. tuberculosis is a highly infectious microbe that is the causative agent of tuberculosis (TB). According to the Centers for Disease Control (CDC), one third of the world’s population is infected with TB. In 2015, there were 1.8 million TB-related deaths worldwide. Tuberculosis affects the lungs and other parts of the body. Most individuals have no symptoms (latent infection), but about 10% progress and experience symptoms which are fatal in half of these individuals.
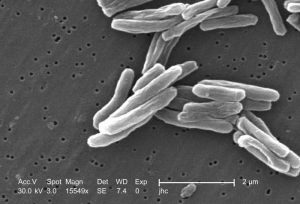
Scanning electron microscopic image of micrograph of Mycobacterium tuberculosis. Photo credit: Janice Haney Carr/CDC
Since the first antibiotic, penicillin, was introduced in 1943, bacteria have evolved over time to resist antibiotics by pumping the antibiotics out through their bacterial cell membrane. Dr. Aiden Doherty, from the University of Sussex, Dr. Jesus Blazquez, from the Centro Nacional de Biotecnologia in Madrid, and their teams used an approach where they knocked-out nearly every gene in mycobacteria and looked at growth of mutant strains in the presence of an antibiotic. The researchers found that the product of the NucS gene, an enzyme involved in DNA repair, significantly reduced mutations.
The NucS gene performs the function of the mismatch repair (MMR) pathway in more advanced bacterial species, which is required for mutation avoidance and anti-recombination. No homologs for the MMR pathway, highly conserved through evolution, had been identified in Archaea and Actinobacteria (to which Mycobacteria belong). Furthermore, the researchers demonstrated that inactivation of nucS in M. smegmatis results in phenotypes that mimic those of canonical MMR-null mutants, including biased mutations, increased mutation rate, biased mutational spectrum and increased homologous recombination.
Antibiotic resistant bacteria is a global health issue
According to the World Health Organization (WHO), addressing antibiotic resistance is a high priority. To that end, the WHO created a global action plan on antimicrobial resistance, including antibiotic resistance, in May of 2015. Drug resistant TB has been found in 105 countries to date.
The United States Centers for Disease Control (CDC) estimates that each year in the U.S. alone, more than two million people are infected by antibiotic-resistant bacteria, leading to at least 23,000 deaths. In 2016, the U.S. Congress appropriated 160 million dollars to the CDC to tackle the crisis.
What can we do to combat antibiotic resistance?
Since the 1940s, antibiotics have been utilized to treat and save many patients from disease. Rampant and inappropriate use of antibiotics is one of the main reasons for the antibiotic-resistance problem we now face. The CDC suggests that up to half of the antibiotics used today for the treatment of humans and animals are unnecessary. Antibiotics should be prescribed only when they are absolutely needed to treat a disease, and the appropriate antibiotic for the disease should be administered. Furthermore, individuals should always complete the entire course of antibiotics prescribed to them by a physician. Only taking a portion of an antibiotic course can help to select for and propagate mildly resistant bacteria, which can then adapt to become much more strongly resistant.
The identification of this key gene by Drs. Doherty and Blazquez is important to learning more about how antibiotic resistant bacteria and “superbugs” (those resistant to multiple antibiotics) develop. According to Dr. Doherty, “The discovery that the NucS gene reduces the rate at which mutations occur in these pathogens is a crucial first step towards identifying the genetic factors that influence the onset of antibiotic-resistance. This will enable scientists and clinicians to screen for strains that are most likely to develop drug-resistance and figure out strategies to tackle this serious threat.”
Do you work in infectious disease research?
Visit our catalog at Kerafast.com for a wide range of products for bacteriology, virology and infectious disease.


