Fighting antibiotic resistance with bacteriophage
A recent study published in the Nature-affiliated journal Scientific Reports describes the use of bacteriophage, or phage, to reverse resistance of the bacteria Pseudomonas aueroginosa to antibiotics. Phage are viruses that attack bacteria and this particular bacteriophage, OMKO1 (family Myoviridae), was found in a pond in East Lyme, Connecticut, not far from lead author Dr. Benjamin Chan’s stomping grounds as a research scientist in the laboratory of Dr. Paul Turner at Yale University.
Although P. aeruginosa can be found in a variety of environments, it has become particularly dangerous over the past few years due to its acquisition of resistance to multiple antibiotics. Individuals with compromised immunity are particularly at risk for infection by P. aeruginosa, which can be acquired anywhere in the community but has proven most deadly when acquired during a hospital stay or at nursing homes, according the Centers for Disease Control (CDC).
Since the first antibiotic, penicillin, was introduced in 1943, bacteria have evolved over time to resist antibiotics by pumping the antibiotics out through their bacterial cell membrane. Dr. Chan and his colleagues found that OMKO1 acts by attaching to the cell membrane and decreasing the efficiency of this pump. Similar to gaining resistance to antibiotics, the Pseudomonas will evolve to resist the OMKO1 bacteriophage. However, in an interesting evolutionary twist, gaining resistance to the phage will then force the Pseudomona to be more susceptible to antibiotics again as the pump will be back to normal function.
Antibiotic resistant bacteria is a global health issue
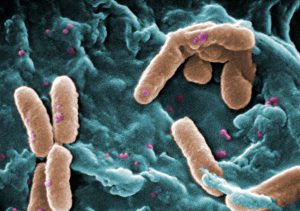
Scanning electron microscope image of Pseudomonas aeruginosa.
Photo credit: Janice Haney Carr, USCDCP
According to the World Health Organization (WHO), addressing antibiotic resistance is a high priority. To that end, the WHO created a global action plan on antimicrobial resistance, including antibiotic resistance, in May of 2015.
The United States Centers for Disease Control (CDC) estimates that each year in the U.S. alone more than 2 million people are infected by antibiotic-resistant bacteria, leading to at least 23,000 deaths. In 2016, the U.S. Congress appropriated 160 million dollars to the CDC to tackle the crisis.
What can we do to combat antibiotic-resistance?
Since the 1940s, antibiotics have been utilized to treat and save many patients from disease. Rampant and inappropriate use of antibiotics is one of the main reasons for the antibiotic-resistance problem we now face. The CDC suggests that up to half of the antibiotics used today for the treatment of humans and animals are unnecessary. Antibiotics should be prescribed only when they are absolutely needed to treat a disease, and the appropriate antibiotic for the disease should be administered. Furthermore, individuals should always complete the entire course of antibiotics prescribed to them by a physician. Only taking a portion of an antibiotic course can help to select for and propagate mildly resistant bacteria, which can then adapt to become much more strongly resistant.
C. Lee Ventola recently published a paper reviewing the current antibiotic resistance crisis and highlighting some of the main contributing factors including the aforementioned inappropriate prescribing, agricultural overuse and lack of regulation – as in some countries antibiotics are available over the counter. He also recounts the regulatory barriers and high expense of creating new antibiotics, which has made them less profitable for pharmaceutical companies and thus a less attractive drug development pipeline target.
Are bacteriophage the answer?
More research is ongoing, but early clinical trial results are promising. According to Dr. Turner, “the virus should help preserve our limited antibiotic arsenal in combating deadly bacteria.” The use of a combined therapy of phage and antibiotics is also a potential treatment for dangerous bacterial infections.
Do you work in infectious disease research?
Visit our catalog at Kerafast.com for a wide range of products for bacteriology, virology and infectious disease.


