A New Avenue for Protein-Nucleic Acid Complex Crystallization
Thomas Hollis, PhD, Wake Forest University — Published: May 22, 2013
The conception of the Protein-Nucleic Acid Complex Crystal Screen came from a need that we had to crystallize DNA-protein complexes. Normally, initial crystallization conditions are found through the use of commercially purchased screens. In the case of DNA-protein complexes, these screens have proven quite unreliable. More often than not, we found that crystals produced using these screens contained protein only or DNA only despite the fact that the protein-DNA complexes used were known to be strongly associated with each other. It seemed that a lot of the conditions in the commercially available screens were actually leading to the dissociation of the protein-DNA complexes.
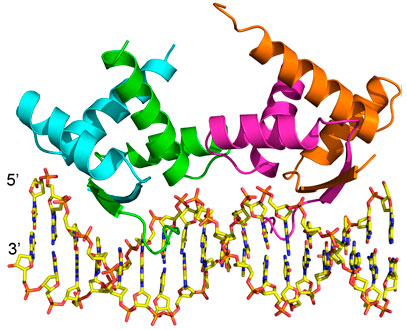
Structural overview of AmrZ – amrZ1 complex (PDB ID: 3QOQ). The Δ42 AmrZ protein binds to the 18 bp amrZ1 binding site as a dimer of dimers. One dimer is composed of chains A and B (green/cyan), while the other dimer is composed of chains C and D (magenta/orange). Adapted from: Pryor EE Jr, Waligora EA, Xu B, Dellos-Nolan S, Wozniak DJ, and Hollis T. The Transcription Factor AmrZ Utilizes Multiple DNA Binding Modes to Recognize Activator and Repressor Sequences of Pseudomonas aeruginosa Virulence Genes. PLoS Pathogens. 2012; 8(4): e1002648.
And so there was a need to develop a screen that would promote the stability of the protein-DNA complex and crystallization. We started by looking through the Biological Macromolecule Crystallization Database protein crystallization database (BMCD) in order to sort out all of the proven crystallization conditions for protein-DNA complexes. Once analyzed, it was very clear that the actual conditions leading to protein-DNA complex crystals came from a relatively small subset of conditions (many of which were not present in commercially available protein-DNA complex screens). One observation was that many of the BMCD conditions contained divalent metal ions. So we thought it might be useful to create our own screen that had all of the conditions in some array form that promote protein-DNA complex crystallization.
We used a technique called incomplete factorial design which has been used previously in crystallography and published on by Charlie Carter at UNC-Chapel Hill. Using the incomplete factorial method in combination with the BMCD conditions that we collected, we designed the screen. This allowed for the thousands of different crystal-producing BMCD conditions to be summarized into just 48.
Basically, the incomplete factorial method enabled us to create a screen that touches a lot of that array space and in different combinations so that almost every element is in combination at least once with one other element in the screen. Therefore, it allows you to sample a very large array space, (in our case the different parameters of PEG concentration and molecular weight, differences in pH, and differences in ionic strengths and divalent metal ions) in a relatively small screen.
We then created the screen and began using it for protein-DNA complex crystallization in our lab. Since it is only 48 conditions, we typically pair it with the PEG-ION screen, which is a predominantly PEG-based crystallization screen. What we have noticed is that we have had tremendous success with obtaining protein-DNA complex crystals especially compared to the commercially available protein-DNA complex screens. It seems that the current commercial screens were not properly targeting the space that promotes protein-DNA crystallization.
Over the past few years we’ve been able to crystallize and determine structures of 10-12 different protein-DNA complexes and every single one of them has produced crystals in this screen. It’s been a tremendous tool. We have published a paper describing the methods for this that came out in Acta Crysallographica in 2012 and it’s been a tremendously useful tool for us in advancing our work. The screen was made available by Kerafast a few years ago and based on feedback that I have received from other researchers, it is a success.
Learn more about the Protein-Nucleic Acid Complex Crystal Screen »
Tags:
Macromolecular cystallography, Protein-nucleic acid complex, Incomplete factorial, AmrZ, Protein-Nucleic Acid Complex Crystal Screen
References:
- Pryor EE Jr, Wozniak DJ, Hollis, T. Crystallization of Pseudomonas aeruginosa AmrZ protein: development of a comprehensive method for obtaining and optimization of protein-DNA crystals. Acta Crystallographica Section F. 2012; F68: 985-993.
- Carter, C.W., Jr., and Carter, C.W. (1979). Protein Crystallization Using Incomplete Factorial Experiments. J Biol Chem 254, 12219-12223.