Exploring the Structure of Human RNA Polymerase I
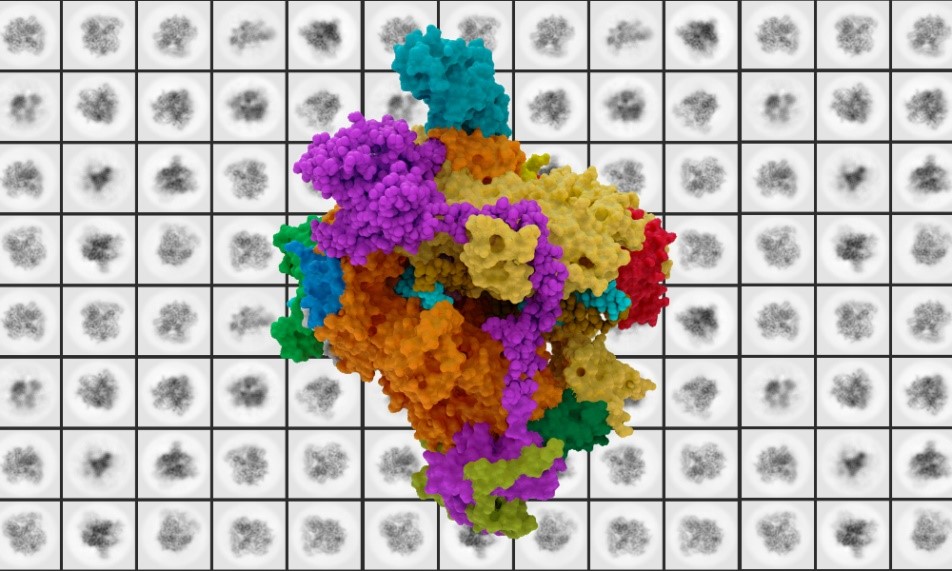
Research led by European Molecular Biology Laboratory (EMBL) scientists has been able to offer a more in-depth look into the connection between RNA polymerase I (Pol I) and mutation-linked diseases. Cryo-electron microscopy enabled researchers to view the structure of human Pol I at a more detailed level than ever before, insight that could be useful for better understanding rare diseases and cancer. Their most recent findings were published in the Nature Structural and Molecular Biology journal.
The study cited our LOBSTR E. coli Expression Strain, which was developed by the lab of Thomas U. Schwartz, PhD from the Massachusetts Institute of Technology for the expression of recombinant polyhistidine-tagged proteins.
RNA Polymerase I (Pol I)
RNA polymerase is a fundamental enzyme in the process of copying DNA into RNA. With three different types, each plays its own distinct role in the transcription process. This study offers a deeper look into the structure of RNA polymerase I, which is responsible for encoding large portions of ribosomal RNA in the body. However, when encoding of these ribosomal RNA strands goes wrong and rRNA synthesis is blocked, cells can run into issues such as apoptosis, senescence or autophagy that may be the cause of a range of rare diseases. It has been found that upregulation of Pol 1 is linked to cancer, while mutations are linked to developmental disorders.
Jumping in Headfirst with Cryo-electron Microscopy
The Müller group at EMBL uses advanced cryo-electron microscopy (cryo-EM) and CRISPR-Cas9 genome editing to explore the structure of RNA Pol I. Their research initially began in 2013 with studying Pol I in yeast, but it was not until recently that they shifted their focus to studying human Pol I. When looking at both the yeast and human Pol I at a microscopic level, they found an important difference in their structures.
On switching to studying human Pol I, lead author Agata Misiaszek said in a press release, “When I started working on this project, we didn’t even know exactly how many subunits comprised human Pol I.”
Their research revealed that yeast Pol I consists of 14 units, while human Pol I consists of only 13. In fact, most species apart from yeast and other fungi have 13 subunits, so scientists were studying an outlier. Upon further examination, the location of this missing subunit is the spot of a critical interaction site, which could be important in understanding human Pol I regulation and its link to rare diseases.
Just the Beginning
Researching human Pol I can be a challenge, as the inability to obtain large amounts at a time makes it difficult to use classic research methods such as X-ray crystallography. However, new techniques such as cryo-electron microscopy and CRISPR continue to emerge, allowing researchers such as the EMBL team to make strides in understanding the interaction of Pol I in human cells.
The LOBSTR E. coli expression strain used in the study is available for researchers worldwide in the Kerafast catalog here. Other related reagents you might be interested in include:
- MarathonRT Reverse Transcriptase from Yale University
- Non-Natural Promoter T7 Polymerase from University of Texas at Austin
- RNA Dependent RNA Polymerase R2 Antibodies from A*STAR Institute
- Protein-Nucleic Acid Complex Crystal Screen from Wake Forest School of Medicine
- DNA Polymerase Eta Antibody from Icahn School Of Medicine at Mount Sinai


